Modelling long-range atmospheric transport of crop diseases
Some of the greatest challenges to food production in low- and middle-income countries arise when crops are devastated by the sudden outbreak of destructive migratory insects and diseases. Pests and pathogenic spores can be channelled by air streams over long distances, crossing international boundaries and continents, bringing severe consequences for crop production and livelihoods in previously uninfected geographical areas.
Forecasting risks of invasion at continental scale, requires simulation models and visualization tools for estimating pest dispersal, range and intensity (spore or insect load), to forewarn national plant health agencies and enable contingency plans to be implemented in advance. The challenge is the quantity of data that are involved for interpretation and visualization of spatiotemporal dynamics of atmospheric transport of crop pathogens and insect pests, requiring tools that are capable of processing the data in a reasonable time, so that pest and disease warnings can be rapidly disseminated to regional plant health authorities, government agencies and extension services, in countries at risk.
Cassava Responsive
Location: Sub-Saharan Africa
Developing a predictive model for an emerging epidemic on cassava in sub‑Saharan Africa
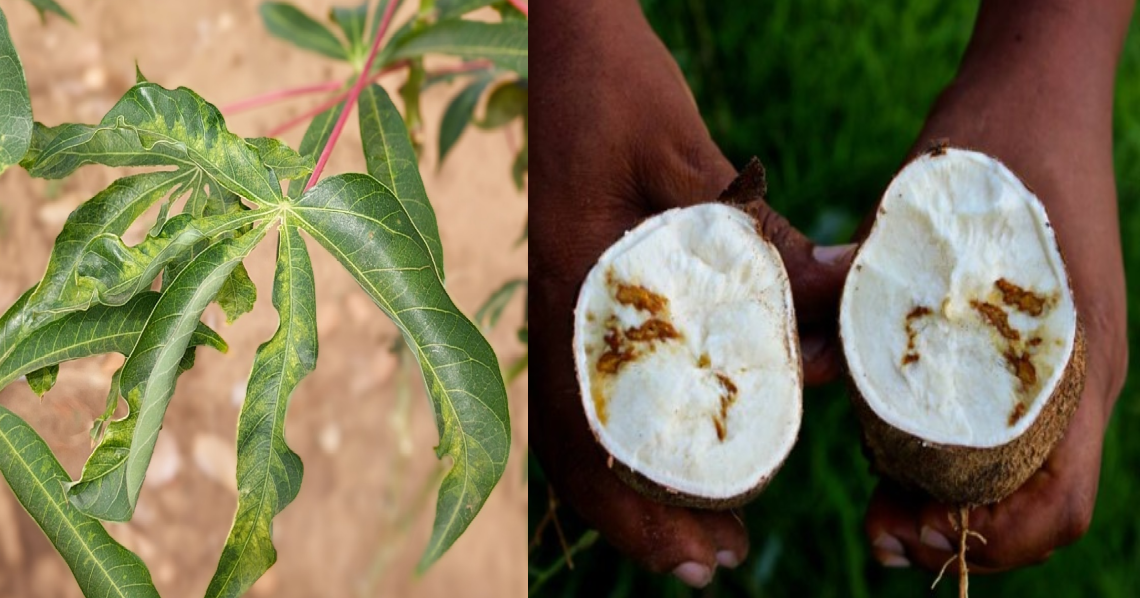

Cassava showing symptoms of cassava brown streak disease in leaves and tubers caused by infections of ipomoviruses.
Cassava (Manihot esculenta) is one of the most important staple food crops produced globally, primarily grown in low-input subsistence farming systems in the tropics and subtropics. In Africa alone, an estimated 800 million people rely on cassava for their primary calorific intake.
Cassava brown streak disease was first reported in Uganda in 2004 but has since been detected widely across countries in East and Central Africa, now reaching as far as the north central districts in the Democratic Republic of the Congo. Continued westward expansion is now a significant threat to food security and economic stability in the major cassava-producing regions of Central and West Africa.
Threatening food production
In recent years cassava production in sub-Saharan Africa has come under increasing pressure due to the rapidly expanding range of cassava brown streak disease (Figure 1). The disease, which causes necrosis of the edible root tissue infected by ipomoviruses, can be transmitted across cassava-growing regions in two ways. Firstly, vectored by Bemisia whitefly, which feed on the aerial tissues of infected plants causing cross contamination within and between field crops. Secondly, through human movement of virus-infected propagated cuttings that are traded, transported and planted for the next season’s harvest.
Predicting epidemics
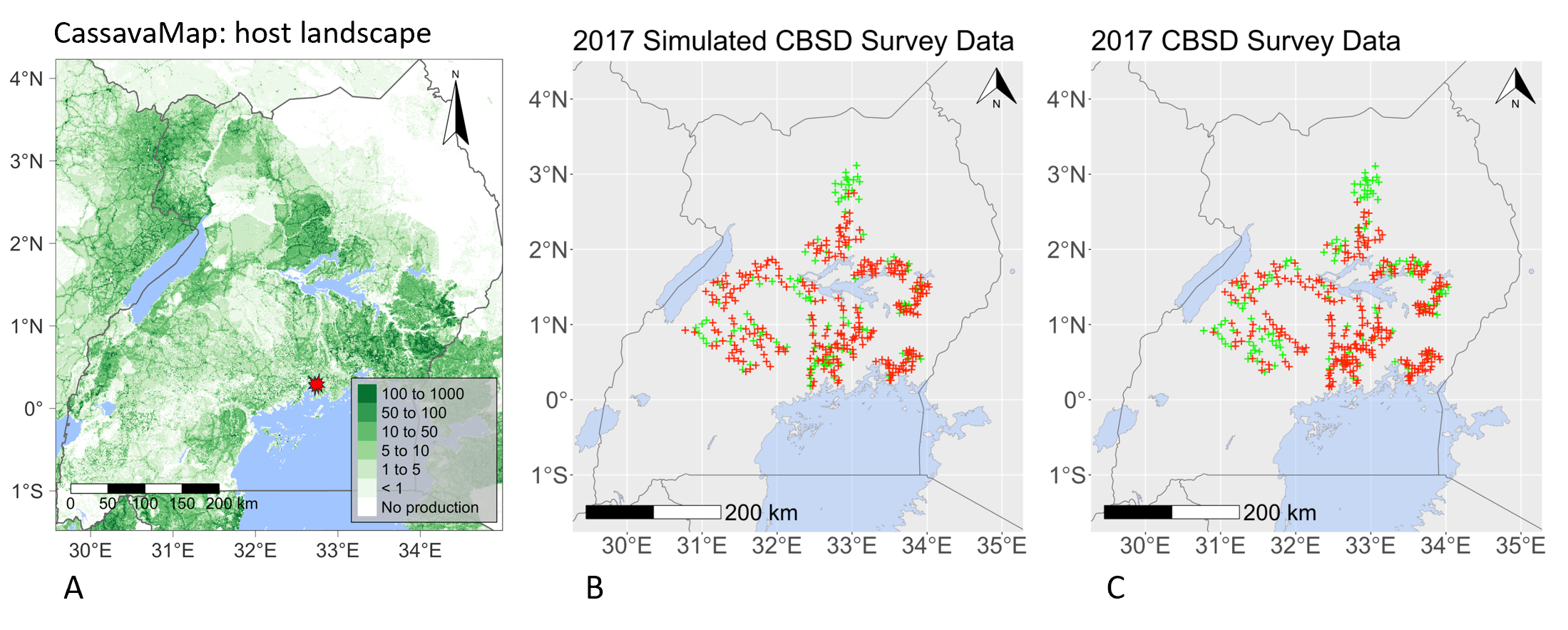
Maps representing the rasterised data-driven model layers: (A) the model cassava landscape, representing the number of fields at a 1 km2 resolution derived from the digital resource, CassavaMap. Testing the model by (B) simulating the spread of cassava brown streak disease using data from 2004 to predict its range in 2017, versus (C) the actual survey data collected in 2017 from the Ugandan field surveys.
Landscape models capable of predicting where and when cassava brown streak disease will occur in a new region are essential tools for ensuring that national strategies can be prepared to mitigate the threat to cassava production areas. Epidemiology and Modelling Group focus on key questions to address challenges faced by national decision makers i) when will the virus cross international borders to arrive in unaffected countries in West Africa; ii) how can these disease-free countries optimise surveillance for early detection and be ready for outbreaks and iii) what strategies can the increasing number of disease-endemic countries use to minimise the impact on farm businesses and food security? Accurately predicting the arrival of a crop pest or pathogen over large regions in sub-Sahara Africa is extremely challenging due to the complexity of crop landscapes within and between countries. In a recent study, members of the Cambridge team used surveillance data for cassava brown streak disease collected by colleagues in Uganda, to design and validate a stochastic epidemic model of the virus, as it spread to new districts. The new model provided a framework for analysing the spread of cassava brown streak disease (Figure above), taking into account the variation in the distribution of the cassava fields and the prevalence of the whitefly vector across the Ugandan landscape.
Developing a model framework for future management strategies
Moreover, the research team designed the model to simulate the impact of planting virus-free cassava cuttings (clean seed programme) in localised areas around Kampala, and were able to demonstrate its use for predicting and analysing the impacts of disease-management scenarios. Gaining an insight into the regions threatened by epidemic outbreaks of cassava brown streak disease is key in supporting the chain of events that will ensure timely guidance and resources are available to agencies and farmers to minimise the risk of crop losses.
[Godding, D., Stutt, R.O.J.H., Alicai, T., Abidrabo, P., Okao‑Okuja, G., Gilligan, C.A. (2023), Developing a predictive model for an emerging epidemic on cassava in sub‑Saharan Africa. Scientific Reports, 13: 12603], (https://doi.org/10.1038/s41598-023-38819-x)
Project members
- Dr Frank Ndjomatchoua
- Dr Rich Stutt
- Dr Ruairi Donnelly
Wheat Rust Early Warning System
Location: Asia & East africa

Wheat rust diseases are caused by airborne fungal spores and pose a major threat to food security all over the globe, especially in countries in East Africa and South Asia. New emerging strains of the disease present an intensifying risk of severe crop loss.
As a result of the relentless efforts of many collaborating organisations (University of Cambridge, Met Office, CIMMYT, EIAR and ATA), an Early Warning System has been established to provide a daily near-real time weeklong forecast of the spread of stem (Puccinia graminis), stripe (Puccinia striiformis), and leaf (Puccinia triticina) rust in Ethiopia. The system has recently been extended to include Kenya in East Africa and Nepal and Bangladesh in South Asia (the latter in collaboration with CIMMYT-Bangladesh, NARC, BWMRI, DHM, DAE, and EMBRAPA).
Here is the schematic summary of the EWS components (see further details in Allen-Sader et al. 2019):
The modelling involves a combination of meteorological and epidemiological models with additional inputs from in-country surveillance coordinated by CIMMYT. The team in Cambridge is responsible for the development and maintenance of the modelling elements of the EWS, including the adaptation of a particle dispersal model (NAME) originally developed by the UK Met Office for rust spores. The UK Met Office also provides seven-day weather forecasts for the target countries to drive the models, giving farmers a three-week window to apply fungicides to prevent rust epidemics. The models are also being used pre-season to assess risks from new races of wheat rust and to inform the selection of crop varieties likely to confer resistance.
Wheat rust predictions across sub-Saharan Africa and South Asia. Many factors must be considered e.g. wind-borne pathogen dispersal
 Example deposition (left column) and environmental suitability (right column) outputs of the model for the East Africa
domain (upper row) and for the Ethiopia subdomain (lower row). Movie below includes the Epidemiological Model outputs
for the Ethiopia subdomain.
Example deposition (left column) and environmental suitability (right column) outputs of the model for the East Africa
domain (upper row) and for the Ethiopia subdomain (lower row). Movie below includes the Epidemiological Model outputs
for the Ethiopia subdomain.
Model outputs for the Ethiopia subdomain.
Our team has explored how 3-D visualization technologies can link time-dependant meteorological data to atmospheric transport models to produce 3-D visual representations of pest and disease movements at regional and continental scales.
Visualising spore release (dispersal) from a detection site in southern Pakistan, showing time-lapse release of simulated 3D spore atmospheric transport dynamics. Meyer et al. (2023) Atmosphere
Recent publications and presentations:
Project members
- Dr Jake Smith
- Dr Tamas Mona
- Lawrence Bower
- Dr Rich Stutt
Funders:
This work is funded by The Bill and Melinda Gates Foundation and UK Foreign, Commonwealth & Development Office for East African part of the project and by the UK Foreign, Commonwealth & Development Office for South Asian, as part of the umbrella project: Asia Regional Resilience to a Changing Climate (ARRCC)).
Locust Responsive



Desert locust (Schistocerca gregaria) is one of the most dangerous migratory pests for millions of subsistence farmers across sub-Saharan Africa, making control a priority for food security in many regions. We developed a novel integrated modelling framework for use in predicting desert locust populations giving government agencies, agronomists and farmers time to prepare. The framework integrates selection of breeding sites, maturation through egg, hopper and adult stages and swarm disperse in search of areas suitable for feeding and breeding. Using a combination of concepts from epidemiological modelling, weather and environment data, together with an atmospheric transport model for swarm movement we provide a tool to forecast short-and long-term swarm movements. As climate change leads to new areas being under threat from desert locust invasions, our approach offers a practical starting point for use in tackling upcoming upsurges.
Below, our research shows that integrated aerobiological epidemiology modelling systems are advantageous for explaining how weather events facilitate relocation of desert locust movement between Yemen over the Gulf of Aden and the East African continent.
Take a look at the articles:
Meyer M., Thurston W., Smith J.W., Schumacher A., Millington S.C., Hodson D.P., Cressman K. and Gilligan C.A. (2023), Three-Dimensional Visualization of Long-Range Atmospheric Transport of Crop Pathogens and Insect Pests. Atmosphere, 14, 910. https://doi.org/10.3390/atmos14060910
Retkute, R.; Hinton, R.G.K.; Cressman, K.; Gilligan, C.A. Regional Differences in Control Operations during the 2019–2021 Desert Locust Upsurge. Agronomy 2021, 11, 2529. https://doi.org/10.3390/agronomy11122529
Project members
- Dr Renata Retkute
Banana Bunchy Top Disease (BBTD) Responsive
Location: Benin, Nigeria, Australia
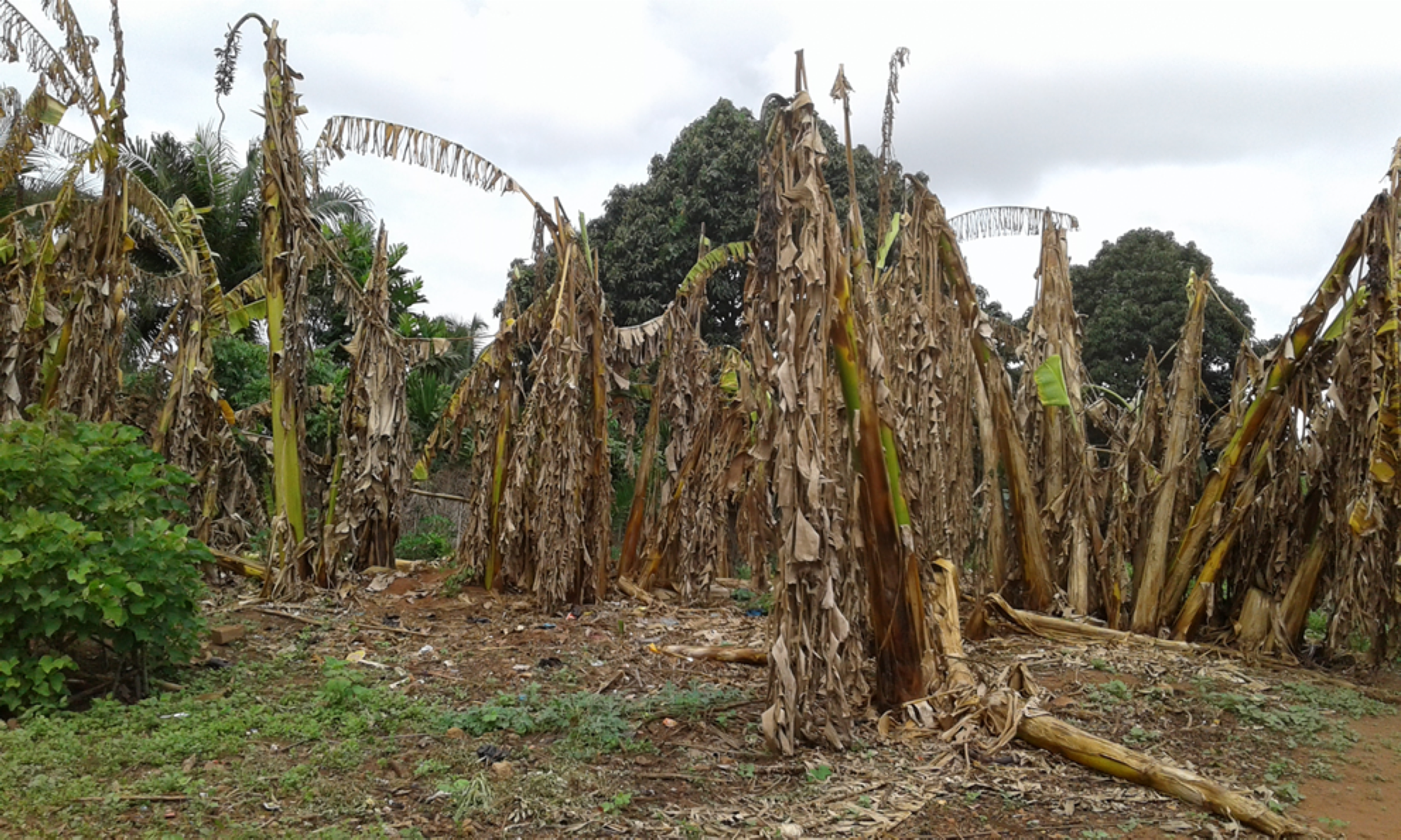

Research focus: Potential of clean-seed programs for re-establishing banana production in disease endemic areas
Investigation of clean-seed program for re-establishing production in disease endemic areas
Despite the increasing use of the dissemination of disease-free planting material, it remains unclear how to optimise the potential of of clean-seed program for re-establishing production in disease endemic areas
Bananas are reintroduced by successively planting banana suckers in 0.5ha fields. Smallholder farmers have sourced clean seed material, which is 100% virus free, to start growing the initial field. All plants produce new suckers from the field are planted to replenish initial field or to expand production into new fields.
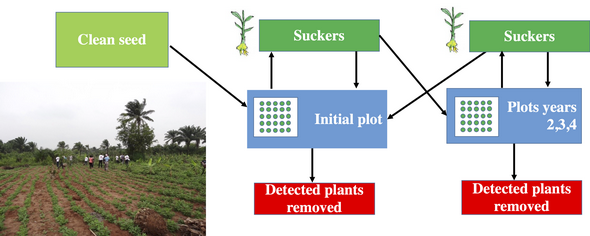
Sustainable strategy
Running generated simulations we can provide evidence that introducing virus-free material in areas affected by BBTV along with propagation of suckers combined with plant inspection for infection and removal of infected plants can provide a sustainable strategy for disease management.
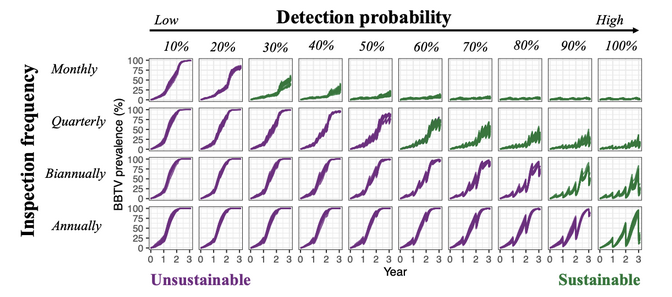
Models were validated using field data from Omondi et al. 2020, Plant Pathology, 69, 1754.
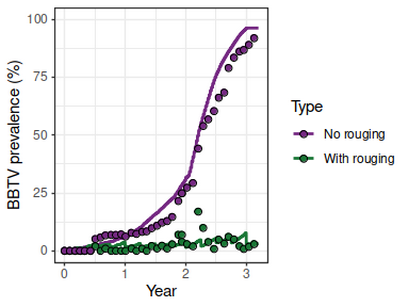
The benefits
This model could be adapted to be used for other hosts and pathogens.
We are able to include additional management options (continuous provision of clean-seed, pesticide treatment) into the model which increases model robustness and accuracy for determining the benefits of introducing clean seed programmes under different farming systems.
Project members
- Dr Renata Retkute
Partners
- University of Queensland, Brisbane, Australia
- CGIAR Bioversity International, Cotonou, Benin
- National Horticultural Research Institute (NIHOR), Nigeria
- Universitate Nationale d'Agriculture (UNA), Benin

